Most people have difficulty comprehending that the universe has 100 billion stars. The fact that the human body harbors 400 times as many microbes is therefore almost inconceivable. Understanding how they function and interact therefore requires very special tools. While discovering and exploring such new biological territories, Professor Peer Bork has developed computer-based tools that analyse the smallest biological entities and that have enabled him but also researchers and companies around the world to develop treatments and environmentally sound production methods. He is being awarded the 2021 Novozymes Prize for this groundbreaking work.
Most people do not associate hype with exact science. Nevertheless, science is often based on hype – popular and interesting theories that unfortunately do not always pass the ultimate scientific test: experimental evidence. Peer Bork has dedicated his career to decode the many experimental data that increasingly flood the world of science today to form hypotheses rather than prove them, which required conceptually new approaches.
“Hype is very common in biology, and some researchers analyse data with a specific hypothesis in mind. I constantly strive to move into some of these new fields and create tools that instead enable data to show us what is actually going on. This has enabled me to create a way of understanding better how microorganisms interact in our bodies, so we can diagnose and cure diseases, and also in the world’s oceans, where this helps us to understand and maintain biodiversity,” explains Peer Bork, Director of the Heidelberg site of the European Molecular Biology Laboratory (EMBL) in Germany.
The easy life in western Germany
Even though Peer Bork has spent most of his 34-year career in Heidelberg, his career actually started somewhere else – in the German Democratic Republic in Leipzig and later on in Berlin, where his PhD supervisor Jens Reich was one of the leading figures in the East German resistance. Doing research sometimes brought about unexpected challenges.
“It was just before the Berlin Wall fell. Jens Reich was under constant surveillance by the Stasi, and he was highly likely to end up in prison. Taking him as a supervisor was therefore risky. Further, if I wanted to read a new scientific article, it took weeks to get it because the library had few journals and we had to beg the authors for reprints, which made things difficult and drawn-out. So my time as a researcher taught me not to take anything for granted,” recalls Peer Bork.
Peer Bork learned from Jens Reich the importance of having a committed and supportive supervisor and of always treating everyone as an equal. But, just as important, his time as a researcher in the German Democratic Republic also taught him that being talented is one thing, but eventually it is commitment that produces great results.
“Life is so easy in western Germany compared with the hard life in eastern Germany back then – or in low-income countries today. So, when I hire PhD students, I do not just choose the best candidates. I want to sense their drive and energy – that they really want to achieve something. So I certainly believe that the lack of certainty in the German Democratic Republic and having to work hard to succeed strongly affected me. Commitment was essential; otherwise you could not achieve anything.”
Similar to LEGO
Peer Bork’s own commitment was sparked by interest in mathematics and computer science in the early 1980s, when he was first exposed to computer programming by practising on an old Russian computer the size of a car that could not properly divide two numbers and needed to be fed by punched tapes. When he was a young MSc student at the University of Leipzig, Peer Bork became fascinated with the potential of solving biological problems computationally, starting with simulating and optimizing the enzyme production processes with differential equation systems. Later on, the enzymes and their functionality inspired him.
“The activity of the enzymes turned out to depend on cofactors – tiny molecules that activate the enzymes. I then became fascinated with the evolutionary development of the areas or domains in the enzymes where the cofactors attach since this is required to understand optimization towards efficient enzyme function," says Peer Bork.
It turned out that the enzymes of interest were all modular, meaning that those cofactor-binding domains were similar across very different enzymes, but this was all insufficiently studied.
"Thus, I started collecting the sequences of these enzymes, focused on the domains that acted like a LEGO block in a more complex structure and tried to find similar domains in new sequence databases," adds Peer Bork.
Since existing software was not sufficient for this, Peer Bork and a colleague programmed a new tool (PAT) based on sequence patterns. With this, he could make his first real biological discoveries showing the widespread nature of those binding domains. The sequence analysis field was still quite small in the late 1980s and early 1990s, but the databases rapidly expanded. Since Peer Bork found biology fascinating and had a natural flair for computers and sequence comparisons, producing another biological innovation was enjoyable and relatively easy.
“I soon turned from just examining the specific predictions of binding sites in enzyme domains to identifying many other types of domains in other types of proteins, since it turned out that this LEGO principle of modularity was very widespread, in particular in extracellular proteins. So I started to collect sequence patterns -“signatures” - of those and to build my first own database with the aim of annotating proteins at that “domain” resolution, knowing that each domain must have a particular subfunction," explains Peer Bork.
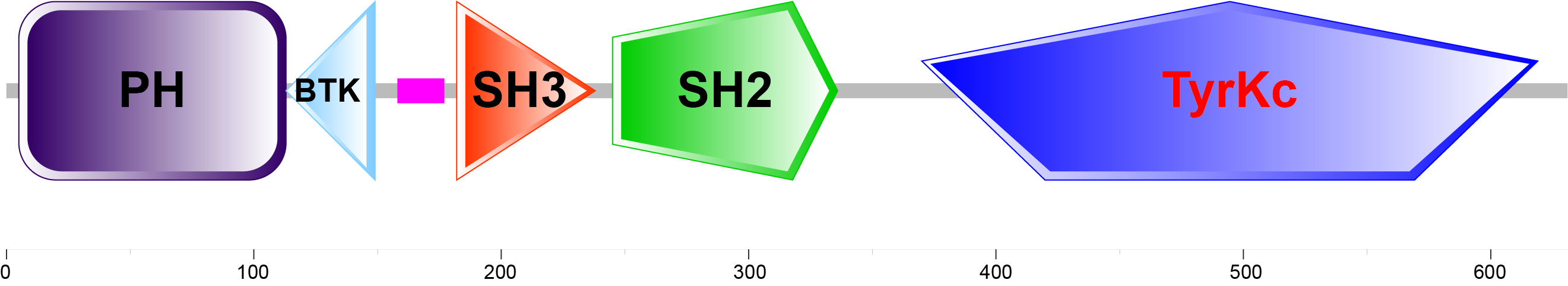
SMART (simple modular architecture research tool) allows rapid identification and annotation of domains in proteins and is able to quickly determine the modular architectures of all gene products in sequenced genomes.
The first smart tool
Like many others in eastern Germany, Peer Bork moved west when the Wall fell. In 1990, having acquired his own funding, he first became a visitor and soon thereafter got a shared appointment at the European Molecular Biology Laboratory (EMBL) in Heidelberg, a university town steeped in tradition in southwestern Germany. EMBL is Europe’s flagship laboratory in molecular biology research, with more than 100 independent research groups at six sites, and the spirit and openness were overwhelming for somebody who had just arrived from the German Democratic Republic.
"The atmosphere was not only multicultural but also extremely collaborative. You could always ask one of your colleagues on another floor to help you. Something that would normally take me 5 years to test by myself suddenly only took a few weeks at EMBL. And because you are only allowed to stay there for a maximum of 9 years, there are always fresh new faces and inspiration, which is great. A continual flow of all these talented and friendly young people with new ideas," recalls Peer Bork.
Peer Bork quickly emerged as a talented young researcher, and his ability to combine computers with biology paid off. In the mid-1990s, he and his colleagues expanded his collection of extracellular domains he brought with him from Berlin and designed a simple but highly effective tool, the Simple Modular Architecture Research Tool (SMART), to predict domains efficiently in the sequence databases that had become much enlarged. This tool was also extremely useful in the early days of sequencing entire genomes, with most of the proteins having no function associated with them, so SMART helped a lot in annotating them.
“By comparing sequences from many classified domains of signalling proteins in SMART, we could quantify their occurrence in entire genomes. For example, we discovered that almost 7% of the genes in the yeast genome contained one or more signalling domains – 350 more than previously estimated. Most importantly, as the World Wide Web had arrived by then, we also build a web resource around this for everybody to use in an intuitive way."
Crucial to biotechnology
SMART was just the first of Peer Bork’s many online tools and was a turning point for him since it was a ticket to get involved in a great breakthrough, not just for himself but also for genetics. Because of his growing reputation, largely because of SMART, he got invited to be part of the most groundbreaking scientific project in the past 50 years: the Human Genome Project.
"I have always been driven to work in new research fields and to help develop the necessary tools. I like participating in collaborative interdisciplinary projects, because this broadens the horizon and implies lots of learning, and they can lead to really great biological discoveries. The Human Genome Project was a good example of that, and SMART became an important tool that helped to annotate the functions of many proteins that the human genome encodes," says Peer Bork.
The experience obtained during the analysis of the human genome enabled Peer Bork to help in annotating most of the early animal genome sequencing projects. Although the genes in a genome can be almost randomly arranged in animals and other eukaryotes, in bacteria the order of genes is greatly constrained: for example, because of bacterial operons, groups of genes that are expressed together to perform a function together.
“The repeated presence of genes in close proximity in the genomes across bacterial and archaeal species indicated a functional interaction between the proteins they encode. As soon as a few bacterial genomes were publicly available, we created the search tool STRING, making use of this observation. With this conceptually novel approach we discovered completely new functionality at a large scale: namely, which interaction partner a protein encoded by a certain gene would have, leading to networks of interacting genes in a genome."
Again, the tool was coupled with a growing resource of already analysed genomes and visuals and made available as a website, using network visualizations that were novel at that time and eventually adding other methods for predicting interacting genes or diverse databases with information on interacting proteins, soon spanning organisms from the entire tree of life, including humans. Using STRING, researchers worldwide explore their own genomic data and deduce the function of proteins and their interdependence.
"As the number of sequences from various organisms increased, researchers and biotechnology companies could thus freely add value to their own data and learn about the functionality of their target organisms or individual proteins. For example, if they wanted to make yeast produce a certain human protein, they could rapidly ensure that not only the individual gene was cloned but also those encoding tightly interacting proteins," explains Peer Bork.

Peer Bork is Director of Scientific Activities at the European Molecular Biology Laboratory (EMBL) in Heidelberg
An interactive tree of life
When analysing the conservation of gene neighbourhoods across a growing number of completely sequenced genomes, Peer Bork and colleagues realized that very few gene neighbourhoods were completely conserved, partly because very few genes were actually present in all organisms, from bacteria to animals. They identified only 40 of these that are present in (almost) all organisms, and only in one copy, implying that they perform the same function.
"With these “marker” genes for an organism, we could now construct the tree of life in a consistent and semiautomatic way, with high accuracy. But we needed a way of displaying various features of this tree, such as annotating the branches with classical taxonomies or adding other various data sets, so the idea of another tool was born: iTOL, the Interactive Tree of Life, for visualizing basic tree information, merging different types of data, customizing the display in various ways and sharing results with others."
Today, the Interactive Tree of Life tool and the associated web resource have more than 20,000 individual users per month and store more than 1.3 million individual trees made by these users to study relationships between genes or organisms from all the kingdoms of life.
"Having an overview of the taxonomy of completely sequenced genomes across the tree of life and how the genes in these interact, the next challenge would be whether one can extend the gene interaction to organismal interactions, like in real ecosystems, given the vast and almost unknown biodiversity on our planet. Instead of simply relying on sequenced individual organisms, we embarked on the daunting task of sequencing communities of different organisms in environmental samples, to catch them in their natural context," says Peer Bork.
The results are metagenomes, which are billions of tiny sequence fragments from a vast number of organisms, most of which were microbial and unknown, so lots of bioinformatics and novel tools were needed to organize these puzzle pieces into genes and organisms. An even bigger challenge was to develop concepts and methods to compare metagenomes.
"Although microorganisms are everywhere, we knew very little about most ambient microorganisms because growing them under standard laboratory conditions is difficult and because they were mixed together, so distinguishing which sequences originated from which organisms and how many came from each one was difficult. With the new methods and analysis concepts developed, we could compare different habitats like sea or soil and could show that the composition of the genes is a molecular fingerprint of an environment," explains Peer Bork.
Gut microbiome divided into three
The ability to reliably quantify the totality of microbes in a sample (the microbiome) by metagenomics led to a kick-start in another habitat, the human body. Peer Bork, together with European colleagues, therefore created the Metagenomics of the Human Intestinal Tract project (MetaHIT). They initially collected and sequenced faecal samples from 124 people from Denmark and Spain – some healthy, some with obesity and some with inflammatory bowel diseases – to determine whether their gut microbes differed.
At that time, people had only just begun talking about the microbiome, and we thought that understanding how intestinal microbes affect human health required being able to compare their genetic fingerprints. But reliably discriminating between health and disease required yet again developing computational tools.These were the basis for several breakthrough discoveries on the human gut microbiome by the METAHIT consortium. After the first basic characterization of a human gut microbiome from a larger group of people in 2010, Peer Bork and his colleagues published an even more remarkable article in Nature the year after, revealing that the people could be divided into three enterotypes, that is gut community composition types, driven by the bacterial genera Bacteroides, Prevotella and Ruminococcus. This story was covered by thousands of newspapers worldwide.
“Although each of us probably has more than 1000 bacterial species in our microbiome, all of us can, like blood groups, basically be divided into these three enterotypes, each with a different community of bacteria. Although we do not know what leads to the enterotypes, they appear to be relatively stable over time: if you have one of these, you likely still have it a year later.”
How medicine affects the microbiome
Studying the human gut microbiome opened up many opportunities and practical applications. Although what is “normal” is still unclear, Peer Bork and colleagues then embarked on finding microbial markers for certain diseases, such as in colon or pancreatic cancer, that turned out to be better than existing cancer markers, with implications for early disease diagnosis.
“After we carved out a microbial signature for colon cancer, we ensured that this signal was robust in different clinical and geographical settings. We examined the faecal metagenomes of almost 400 healthy people and 400 people with colon cancer and found a stable signal that complements the existing non-invasive faecal occult blood tests. So combining the two approaches leads to improved results, which should be translatable into a non-invasive and cheap early screening test.”
The researchers have already patented the approach and the microbial colon cancer markers, developed such a test themselves and are now performing pilot experiments together with a large diagnostics company. They hope that the company will roll out an improved early detection test for colon cancer very soon. A similar project is also underway to developing non-invasive early diagnosis of pancreatic cancer.
Peer Bork and colleagues also found that therapeutic drugs strongly influence the gut microbiome, often more than the disease. In fact, more than one quarter of all drugs directed to the human body affect the gut microbiome.
“Again, this meant developing tools to disentangle the effects of drugs and those of disease. We surprisingly found that different types of medicine affect the individual’s microbiome in very different ways, so even two people with comparable profiles in their microbial species composition probably have unique microbial features that respond differently to the same medicine.”
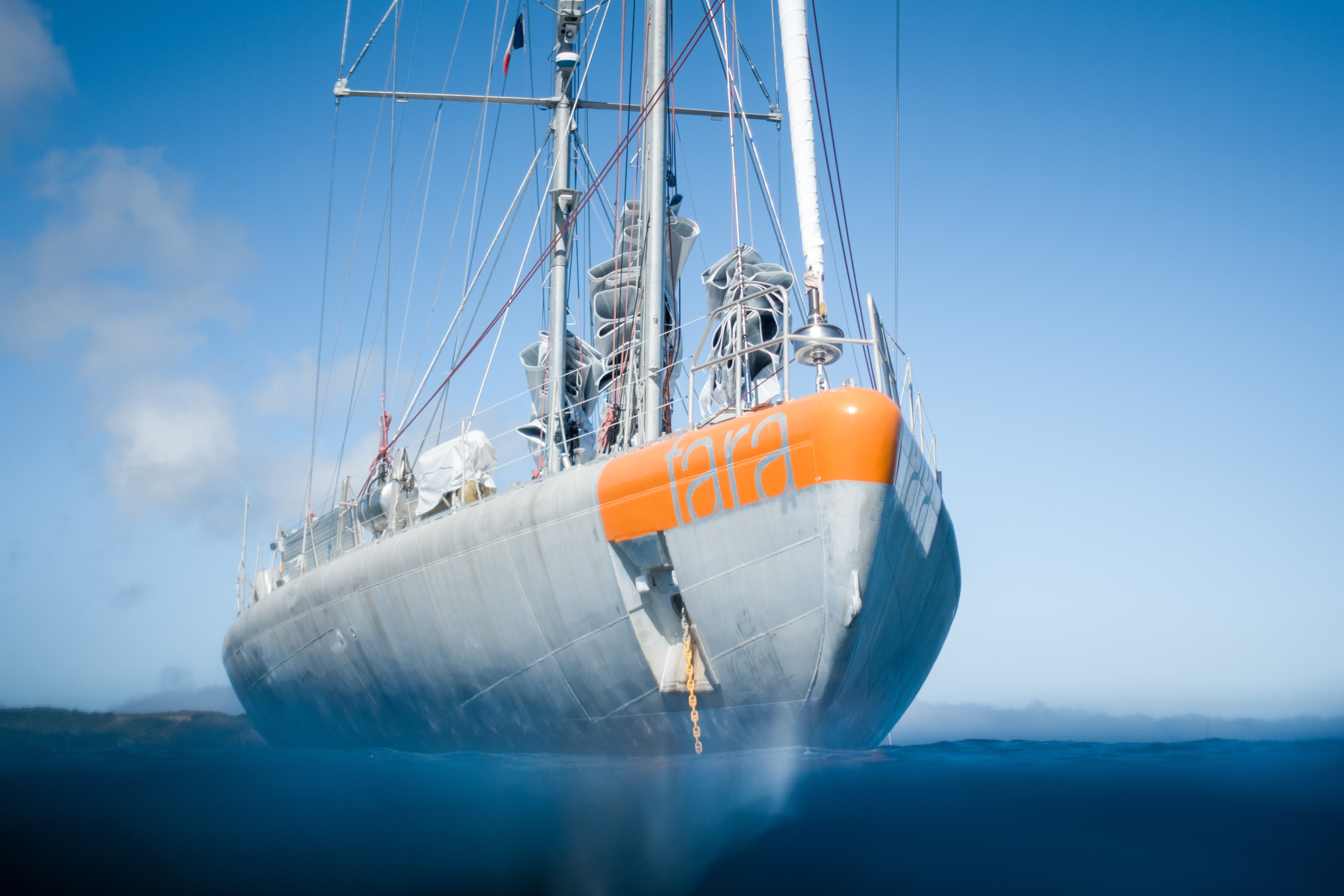

The schooner Tara has been used for a number of global expeditions including ones into the Arctic and Antarctic. A large research consortium (Tara Oceans) is commited to analyse the collected samples. The expeditions also included training programmes and outreach activities to raise public awareness of environmental and climate protection.
The planet's microbiome?
They are currently studying, also using their resource STITCH on chemical–protein interactions, how this knowledge can be used to optimize drug treatment for each person individually after surveying their gut microbiome. Like in the field around the human microbiome, Peer Bork made many basic science discoveries and attempted to translate them into practice, illustrated by cofounding five biotech companies.
With more than 650 scientific articles, 260,000 citations and an h-index of 214, he is unequivocally a leading researcher in the life sciences. Nevertheless, he is far from finished making great discoveries. Instead, Peer Bork and his team continue to plunge into brand new fields, thereby always needing to develop tools that he then shares, also to benefit researchers in biotechnology.
“Our goal with microbiome research has been to improve people’s health. As a bioinformatician, complex samples come to you as digits, 1s and 0s, so it is easy to think about analogies, and using the microbiome for people’s health can in principle also be applied to approach the health of the planet. What if we could study the planet’s microbiome in the same way we study the human microbiome? Microbes can serve as early markers for pollution or unhealthy ecosystems analogous to cancer biomarkers, and perhaps they can be used at some point to remediate ecosystems that are in poor shape, analogous to disease therapy.”
This all requires understanding biodiversity, and very little is known about microbes. Peer Bork turned recently to a microbial census of our planet. For example, as part of the Tara Oceans Consortium around a research sailing boat, which collects plankton around the world at depths down to 1,000 metres, he and his colleagues catalogue and analyse microbial species and their genes in the world’s oceans.
"We have now created a microbial reference catalogue for the oceans that encompasses plankton diversity, viruses, prokaryotes and eukaryotes. We have also developed ways of studying interactions between them and could quantify how environmental factors such as temperature affect ocean microbial communities," says Peer Bork.
The tools must be good enough
Peer Bork believes in creating much better evolutionary understanding of the dynamics of the marine microbial communities, especially given climate change. Therefore, they have collected and shared all the molecular, morphological and environmental data. This has really expanded their understanding of the microbial ecology of the ocean.
"We can now understand and assess the current changes in this ecosystem and hopefully can regenerate it, thus helping to secure the future habitability of our planet. We also analysed topsoil around the world, discovering a global war between fungi and bacteria, with antibiotics and antibiotic resistance genes as weapons and defence systems, respectively. So we can compare microbial communities in different habitats to study how individual genes and entire species evolve and how they spread across the globe."
Peer Bork's journey from working with very small protein domains, large genome sequencing projects and his studies of the human, ocean and, most recently, the Earth’s microbiome, has made him one of the best known and most frequently cited researchers in the world. In fact, almost everyone who analyses genome or metagenome data has used a method or tools developed or influenced by Peer Bork. This really motivates this great scientific toolmaker.
"You always make a special effort to ensure that other people can actually use what you develop. The tools must be good enough to analyse the data, but they must also be intuitive and easy to use. If you have spent time creating a tool and have shared it with others, then knowing that other people actually use it and share their own data is very satisfying, so we can help each other to create a future with a healthier global population and a more sustainable planet," concludes Peer Bork.
